Discrepancy Results JASP & BayesFactor (R)
Hello,
I am doing a Bayesian RM-ANOVA with both JASP and the BayesFactor package in R. I am using the example data set - Bugs (Ryan, Wilde & Crist, 2013) - for RM-ANOVA, provided by JASP. I am only using 2 (Disgusting) X 2 (Frightening) as the repeated measures variables. For some reason I get rather different results in JASP and in R.
In JASP I kept the default settings:
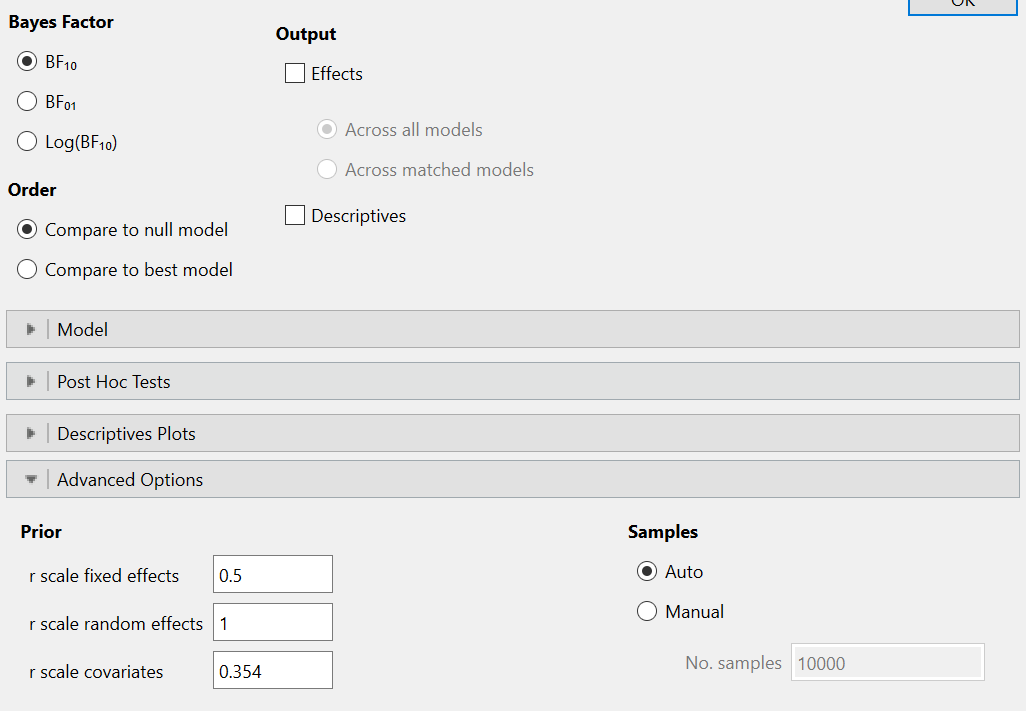
I get the following output:
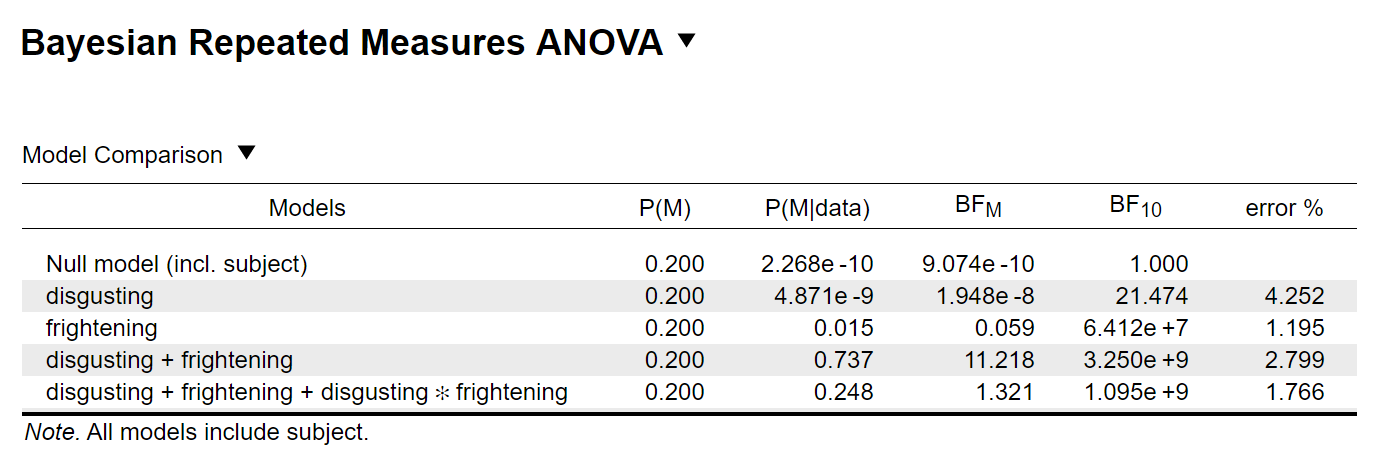
In R, I first put the data into the long format and excluded missing values (because BayesFactor throws an error otherwise). You can see the script here:
library(tidyverse)
library(BayesFactor)
data <- read_csv(file = "Bugs (Ryan, Wilde & Crist, 2013).csv")
data_cl <- data %>%
gather(key = "key", value = "Value", 5:8) %>%
separate(col = "key", into = c("Disgusting", "Frightening"), sep = ", ") %>%
filter(!(is.na(Value))) %>%
mutate_at(c("Subject", "Disgusting", "Frightening"), as.factor) %>%
as.data.frame()
mod_bay <- anovaBF(formula = Value ~ Disgusting * Frightening + Subject,
data = data_cl,
whichRandom = "Subject")
For the R analysis I get the following results:

I am aware that the results are always a bit different due to Monte Carlo sampling. However, I do not think that this explains such large discrepancies in the Bayes factors between JASP and R.
Does someone have an idea how this comes about?
Thank you very much for you help!








Comments
JASP ought to use the BayesFactor code. My first suggestion would be to run the analysis multiple times to get an indication of the variability in the outcomes. How systematic is it?
E.J.
Thanks for your reply!
Yes, I had the same suspicion. Looking only at the effect of "Disgustin", for example, JASP gives BF10s roughly between 20 and 22 whereas BayesFactor gives BF10s roughly between 30 and 32 when I run the analyses 5 times.
OK. And you checked that it does not have anything to do with the missing values? So when you apply JASP to the data after removing the missing values, the results are unaffected?
E.J.
Well, indeed that is the problem! When I removed the missing values in R, only one row in the long data format (i.e., one cell in the wide format) was removed, whereas JASP seemed to remove a whole row in the wide format. So, adjusting for this, I now get the same results in JASP and R.
Thank you for dissolving that riddle for me!
M.L.