Interpreting results of one-way bayesian ANOVA in JASP
Dear forum,
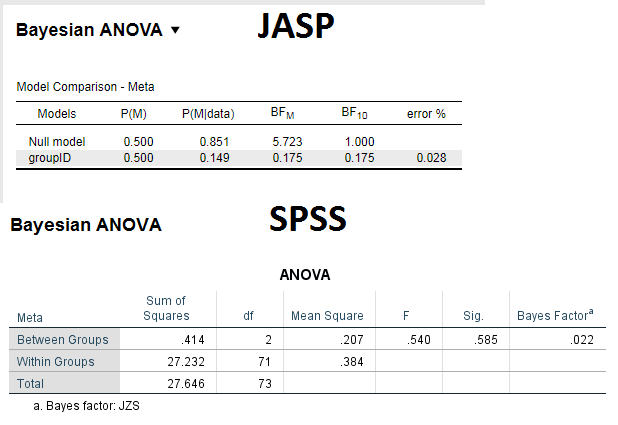
I would like to ensure that I correctly interpret and report the results of one-way bayesian ANOVA (different samples, not repeated measures). I attach an output of my analysis (using JASP and SPSS). My questions are:
1. If I read the output correctly, in JASP I get Bayes factor (BF10) 0.175. Given that it is < 0.33, can I say that this supports H0?
2. When I report my JASP analysis I say that the priors were based on cauchy distribution. This is what I understood from this paper: https://link.springer.com/article/10.3758/s13423-017-1323-7 Would it be correct statement?
3. For comparison, I ran the same analysis in SPSS (Jeffreys–Zellner–Siow method). in SPSS the Bayes factor is 0.022. Is the discrepancy between two programs because of different priors?
Thank you a lot,
Vadim


Comments
Dear Vadim,
Cheers,
E.J.
SPSS is in error, almost surely. It would require a prior scale of almost two to get that low a Bayes factor.
Here's my check. This isn't exactly right, because it appears you have slightly different numbers in each group, but this is the only way I can just use the F statistic. It should be very close.
Thank you very much for your answers and great software!
Hello everyone,
First, many thanks to the JASP team for the amazing job! I have the following naive question
For a study we did 4 one-way Bayesian ANOVAs (for 4 different tasks) for both Reaction times and Accuracies. For the Acc the BF10 and 01 are easy to interpret (meaning that there is clear support either for BF10 or BF01), but for the RTs both the BF01 and BF10 are negligible (less than 1). So my question is: when both BFs are less than 1 would it be correct statement to say that the data (in this case the RT data) is not informative ( probably more evidence is needed) and thus no reliable conclusion can be provided by from this data? Or some other way to handle such small BF10 and BF01?
Thank you in advance!
Hi Mila,
BF10 = 1/BF01, so if BF10 < 1 then BF01 > 1. In other words, it is impossible that both BF10 and BF01 are lower than 1, unless you are referring to different tests. Maybe you have a concrete example? In general, BFs near 1 are not diagnostic, at least not as far as the hypotheses are concerned that you are testing.
Cheers,
E.J.
Thanks for the response. Indeed, you are right I dis not express muself correctly. What I was wondering was wheter for cases like BF10 = 0.956 and BF01 = 1.046, for example, where both BFs are not diagnostic, as you said, would be sufficient to say that the data is not sensitive with respect to hypotheses tested and refrain from any conclusions?
Thanks again!
Hi Mila,
Yes. The correct conclusion is that the data do not provide information: the hypotheses under consideration predicted the data about equally well. Of course, the data may be informative in other ways; for instance, perhaps the posterior distribution is relatively peaked, or perhaps most posterior mass is on positive values of effect size, etc.
Cheers,
E.J.
Many thanks!